Research
Polymer-based Gene Delivery
What we do...
Initially funded by the Carl Zeiss Foundation, our junior research group (Jena Center for Soft Matter (JCSM)) was installed in 2013. It is currently funded as an interdisciplinary BMBF junior research group (NanoMatFutur, project: PolyBioMik) at the JCSM.
We focus on the potential of complex polymeric nanostructures such as micelles, nanogels, and compartmentalized particles for gene transfer in eukaryotic cells. The characterization of polymers assembled nanocarriers and biological processes provides our basis for identifying structure-property relationships.
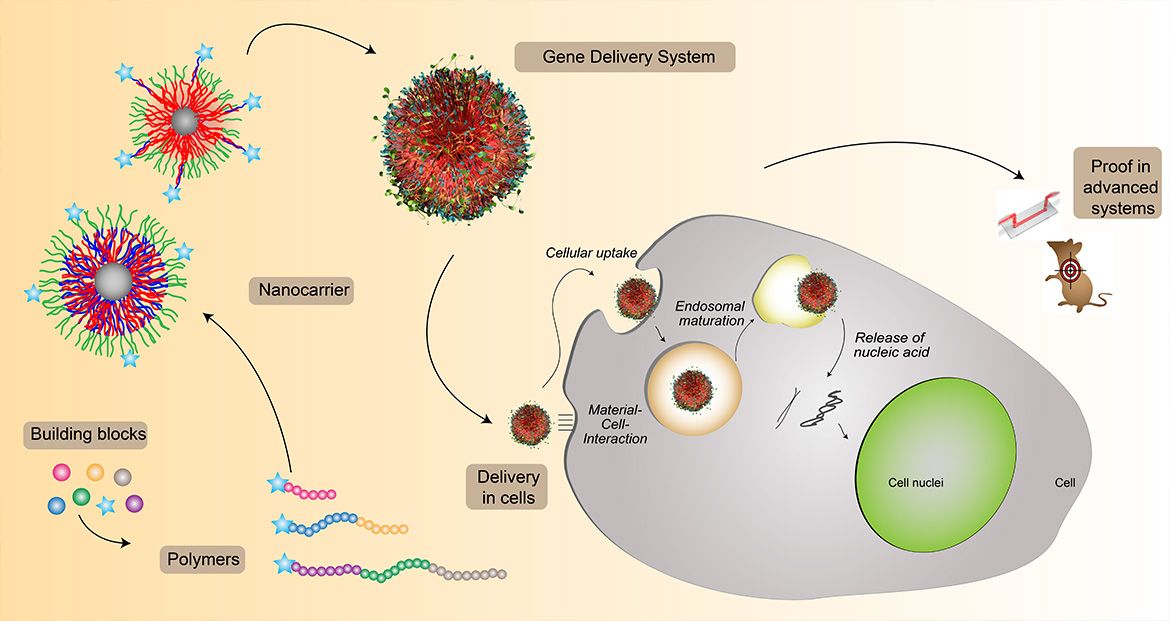
The controlled transport of genetic material or proteins in eukaryotic cells is essential for the development of new therapeutics. Cationic polymers are able to complex the negatively charged nucleic acids to promote cell uptake by electrostatic interactions. When taken up by endosomal processes, the complexes must reach their site of action, the cytoplasm or the cell nuclei. Therefore, the endosomal membrane, an important intracellular hurdle, must be crossed. Here we develop polymers in which different strategies for successful gene transfer by efficient endosomal release mechanisms are used: (I) Polymeric micelles of block copolymers, (II) pH-independent and dependent amine functionalities, (III) integration of hydrophobic units into the polymers.
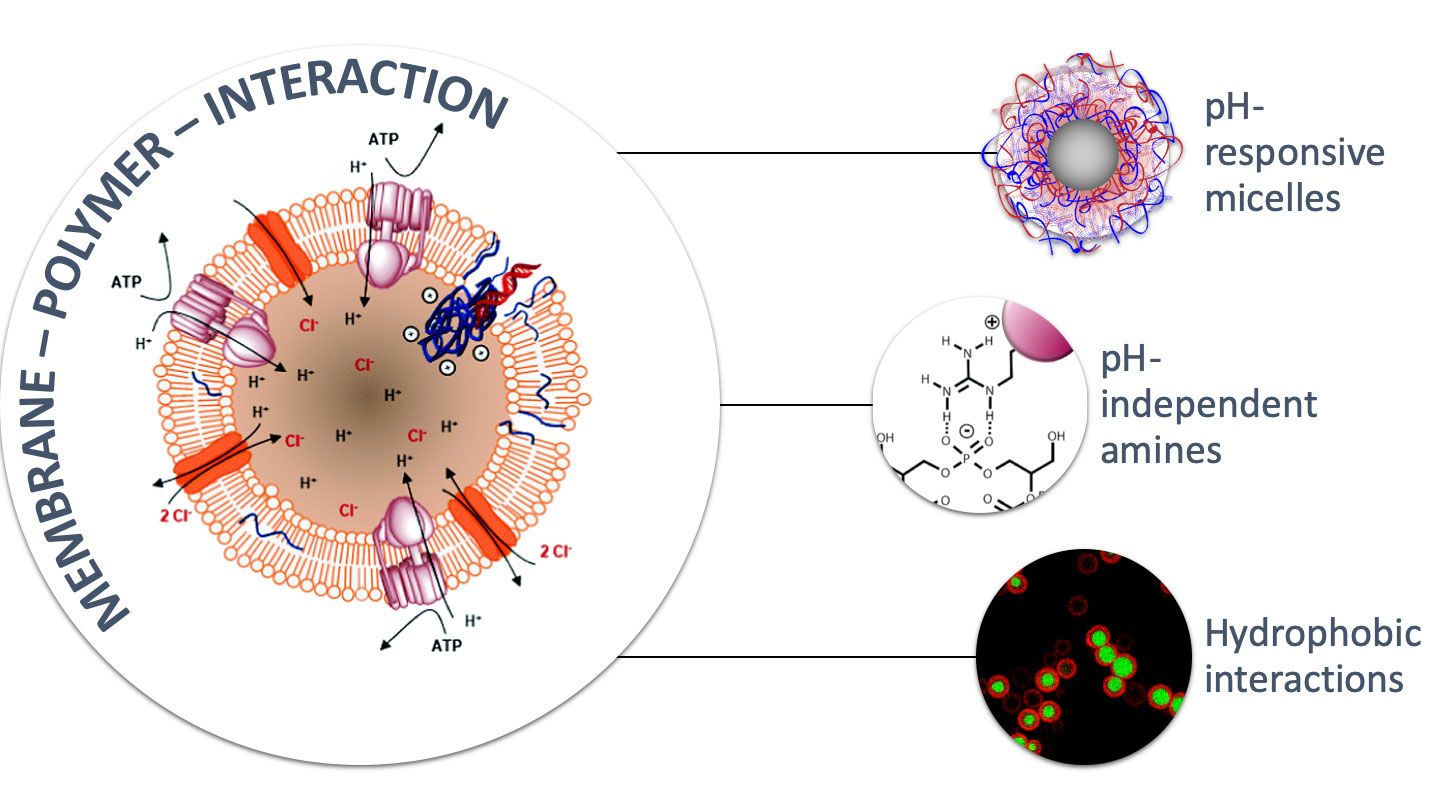
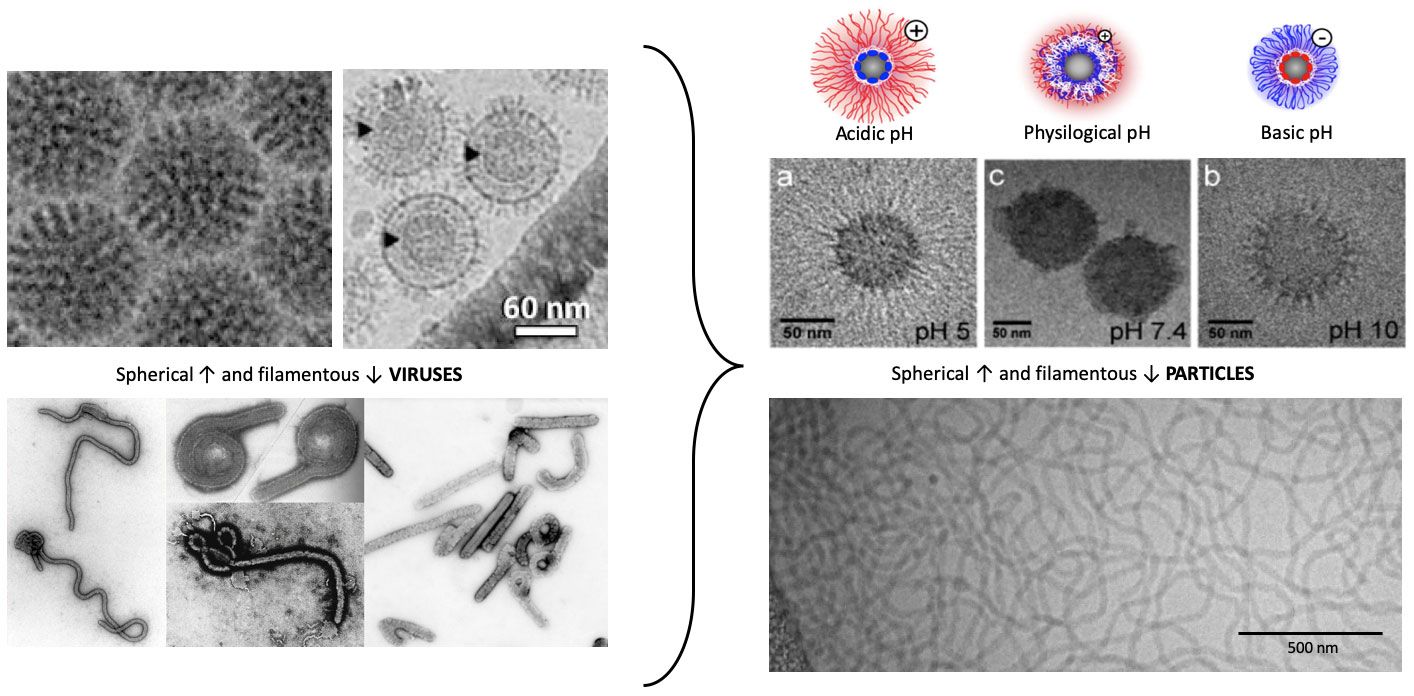
The developed gene carrier are inspired by mechanisms of nature, for example functional groups or structures used by viruses or bacteria. Interdisciplinarity is therefore the focus and is actively lived in our group. Regarding the polymers, we are currently focusing on block copolymers that have different functionalities in the side chains and polymer ends. These polymer structures can assemble into certain defined structures. By this approach, we have been able to increase the efficiency of polymers. We are also investigating the influence of certain functional groups known from nature and present in viral proteins.

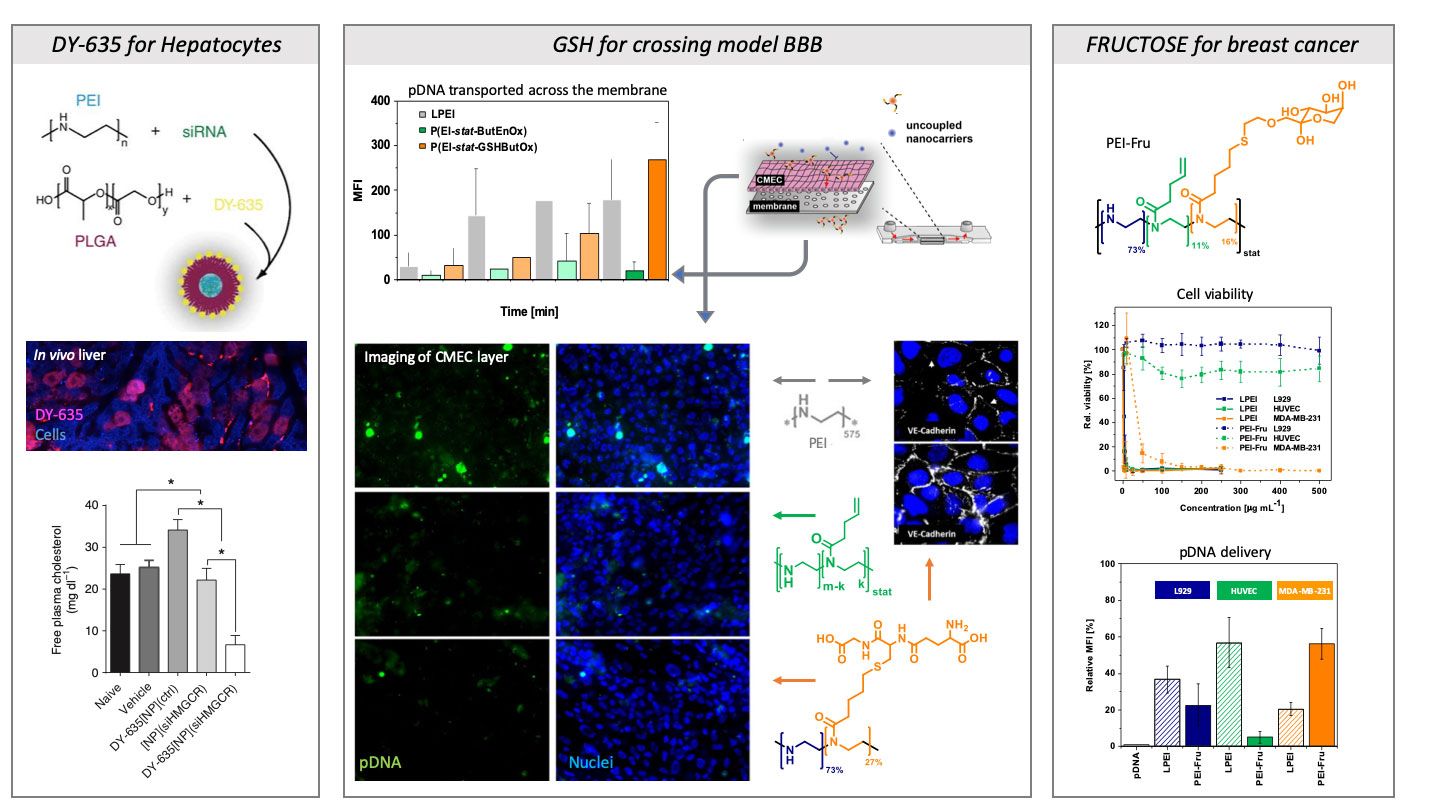
Fokusing on the development of gene carrier for in vivo applications, the delivery to the targeted cell or organ shouis addressed. Therefore, we use so-called target structures that can be coupled to the polymers. Suitable molecules for this purpose are, for example, DY-635, which is a liver-specific dye specifically interacting with the organic anion transporters of the liver (e.g. OATP1B1). With this strategy different drugs can be delivered (further development by SmartDyeLiery GmbH). Nutrients such as fructose can also be used as a targeting unit and, coupled to cationic polymers, could transport genetic material into the triple-negative breast cancer cell line MDA-MB 231.
The blood-brain barrier is known for its restrictive selection and is considered one of the most difficult barriers to overcome in vivo. We were able to couple glutathione (GSH) to polymers, resulting in a significantly improved transfer of genetic material across a modelled barrier in the chip model.
For further information or if you are interested in our research, please contact us.
Our Methods
DNA-Polymer-Interaction
To study the efficiency of the binding between polymeric nanostructure and genetic material, we use the DNA intercalating dye ethidium bromide (EtBr). When mixed with DNA, EtBr shows a high fluorescence intensity due to the intercalation into the double strand. Upon adding a polymer interacting with the DNA, the EtBr is displaced and therefore the fluorescence intensity decreases. These differences in fluorescence intensity can be measured with a microplate reader at appropriate wavelengths. By comparing the results to the positive control of only DNA and EtBr, we can finally determine the relative amount of bound DNA.
For the determination of the stability of the polymer-DNA-complex, we add heparin, a polysaccharide from the extracellular matrix. The polyanion competes with the DNA for the binding of the polymer and finally displaces the DNA from the complex. Thus, EtBr can reintercalate into the DNA, resulting in an increase in fluorescence intensity. High heparin concentrations indicate a strong electrostatic interaction between polymer and DNA.
Internalization of Nucleic Acids
The internalization of nucleic acids is a critical step for efficient gene delivery. To determine the uptake efficiency, cells are incubated with complexes of our polymers and plasmid DNA encoding for a non-fluorescent protein. Before complex formation, the pDNA is labeled with YOYO-1, a green fluorescent, dimeric cyanine nucleic acid stain. Following incubation for different time periods, cells are detached and analyzed via flow cytometry. To make sure, only internalized complexes are measured, the fluorescence of complexes or YOYO-1 labeled pDNA sticking to the cellular membrane is quenched with trypan blue. Again, only viable, single cells are considered for fluorescence analysis. The results consist of two values: i) the amount of YOYOTM-1 positive cell determined by adjusting the gate in the green fluorescence channel to an unstained control and ii) the increase in fluorescence inside the cells determined as the mean fluorescence intensity of all viable, single cells relative to the unstained control (rMFI). Since all cells take up some dye very quickly, the rMFI is the decisive result in this assay.
Endosomal Release
For successful gene delivery, the endosomal escape is very decisive. To determine the efficiency of our nanomaterials in this process, we use the calcein release assay. Calcein is a green-fluorescent, non-permeable dye which is taken up via endocytosis resulting in a punctuate pattern inside the cytosol of cells. If a polymer is able to escape the endolysosome by disrupting the the endolysosomal membrane, this pattern will be changed to a diffuse fluorescence throughout the whole cell. This difference can be observed via confocal laser scanning microscopy (CLSM) and flow cytometry.
Gene Expression
The aim of our research group is to develop efficient nanotransporters for the delivery of genetic material. To determine the transfection efficiency of our in-house designed and synthesized polymers, we use a plasmid DNA (pDNA) encoding for the EGFP protein as a reporter gene. If the pDNA is successfully transported into the cell, it needs to get to the nucleus, where the EGFP-gene will be transcribed with the cell’s own transcription machinery.
The cells are incubated with complexes of our polymers and the EGFP-pDNA for usually 24 h. Following detachment of the cells, the amount of green-fluorescent, EGFP-expressing cells is then determined via flow cytometry. The flow cytometer focuses the cell suspension hydrodynamically, separating the cells from each other to analyze them individually regarding their scattered light. With the help of the forward/sideward scatter plot, only viable single cells are considered for fluorescence analysis. Finally, the results consist of two values: i) amount of EGFP-positive cells (in %) determined by adjusting the gate in the green fluorescence channel to an unstained control and ii) the increase in fluorescence determined as the mean fluorescence intensity of all viable, single cells relative to the unstained control (rMFI).
Assembly of Polymer with Nucleic Acids
For an efficient polymer-DNA-complex formation not only the concentration of both materials is important, but also the amine to phosphate ratio (N/P ratio). We define it as the ratio of polymeric, binding-active amines to the phosphates of the genetic material’s backbone. So far, we showed N/P ratios higher than 1 to be beneficial for efficient gene delivery.
To obtain the polymer-DNA-complexes, we prepare stock solutions of both materials with appropriate concentrations, which are then mixed at a 1:1 ratio, vortexed vigorously and incubated for 15 min. We usually keep the concentration of the DNA stock solution constant.
Intracellular Distribution
Another way to study the mechanism for the transfection efficiency of our nanomaterials is to determine the intracellular localization pattern. Therefore, we stain the cells with different organelle markers, e.g. endolysosomes with LysoTrackerTMRed, and incubate them with complexes of polymer and YOYOTM-1 labeled DNA. Following incubation for different time periods, the cells are analyzed with confocal laser scanning microscopy (CLSM). Since CLSM allows for illumination of only one point of the sample within the focal plane and not the whole sample, the colocalization of different objects in one focal plane can be determined. If endolysosomes and nanotransporters are stained with red and green, respectively, a red and green pattern will be observed inside the cells. Overlapping, i.e. colocalized objects will be depicted as yellow and indicate the localization of our nanocarriers in endolysosomes, whereas only green objects indicate escaping nanocarrieres inside the cytosol. This process can also be applied to other cellular organelles.